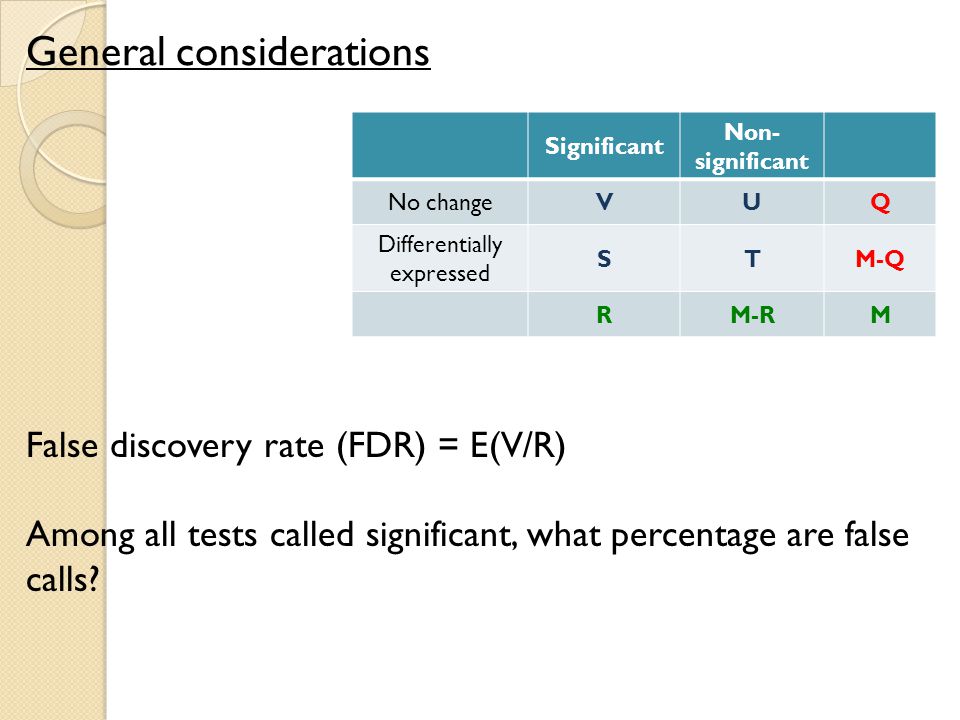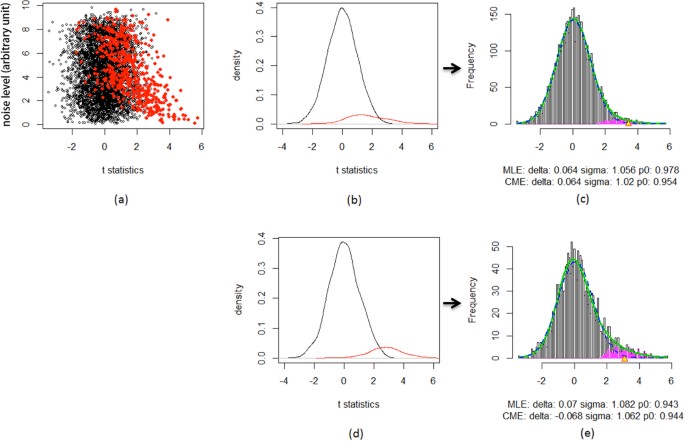
Local false discovery rate estimation using feature reliability in LC/MS metabolomics data | Scientific Reports

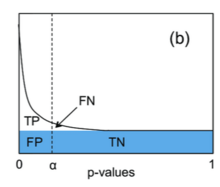
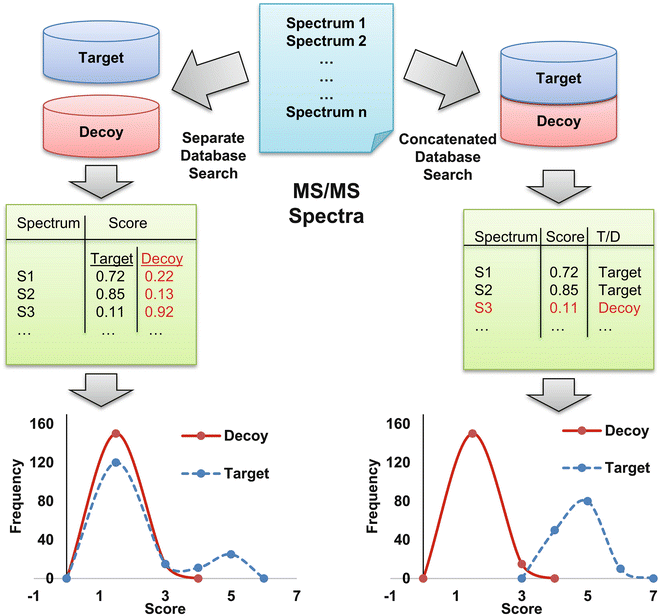
Oliver M. Bernhardt on Twitter: "In this type of FDR calculation, the decoys are only used to estimate the shape and position of your target peptide null-set (non present peptides). The cutoff

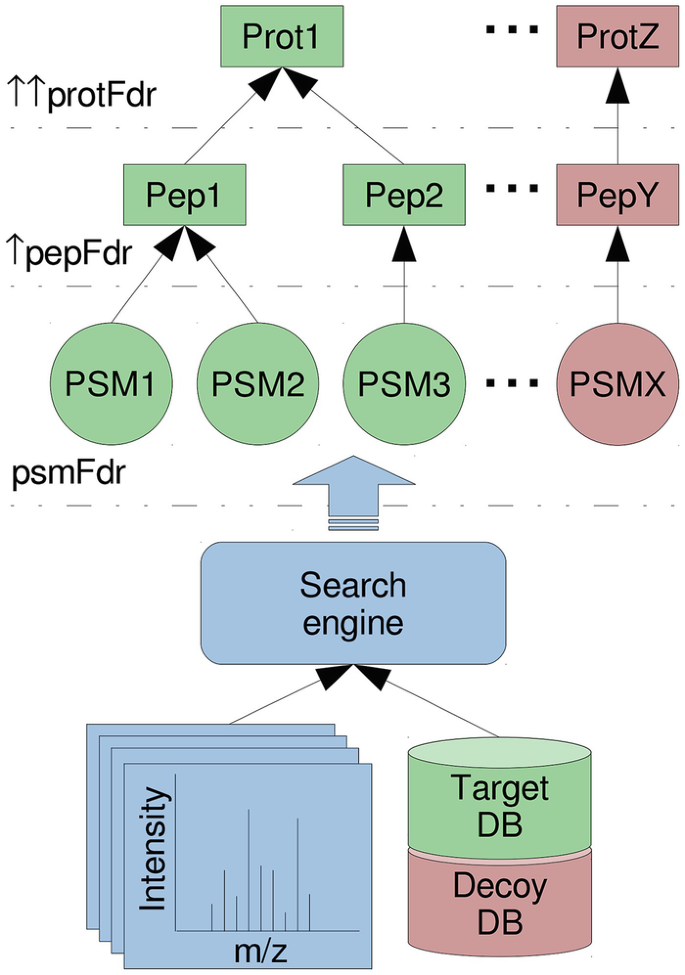
A Scalable Approach for Protein False Discovery Rate Estimation in Large Proteomic Data Sets - ScienceDirect

Local false discovery rate estimation using feature reliability in LC/MS metabolomics data | Scientific Reports